BioUnfold #20 — Transcriptomics: The Control Surface of the Cell
Why measuring regulation can be more powerful than measuring execution.
Transcriptomics became central to modern discovery not simply because it measures many genes at once, but because gene expression is produced by regulatory systems that already constrain what can vary. Transcriptional programs are coordinated and partially redundant. They move within biological limits set by regulation. This makes large-scale integration and statistical modeling far more tractable than it might appear. Other biological readouts — such as morphology, proteomic composition, or metabolic activity — often reflect many overlapping processes without the same governing structure, making them harder to align across experiments. Transcriptomics is powerful because it measures biology in a space that cells themselves have already regularized.
Measurement aligned with regulation
Gene expression is not an arbitrary snapshot. It is the output of networks evolved to maintain stability while allowing controlled adaptation. Transcription factors act combinatorially, pathways reinforce one another, and feedback loops limit extremes. Distinct molecular events often converge on similar transcriptional adjustments because cells respond through coordinated programs.
Transcriptional programs are coordinated and partially redundant. They move within biological limits set by regulation. That bounded variation makes large-scale integration and statistical modeling far more tractable than it might appear.
What RNA sequencing actually captures
In practice, RNA sequencing largely measures messenger RNA, the layer closest to regulatory control rather than biochemical execution. Proteins, metabolites, and morphology realize function but are further shaped by translation, turnover, localization, and chemical context.
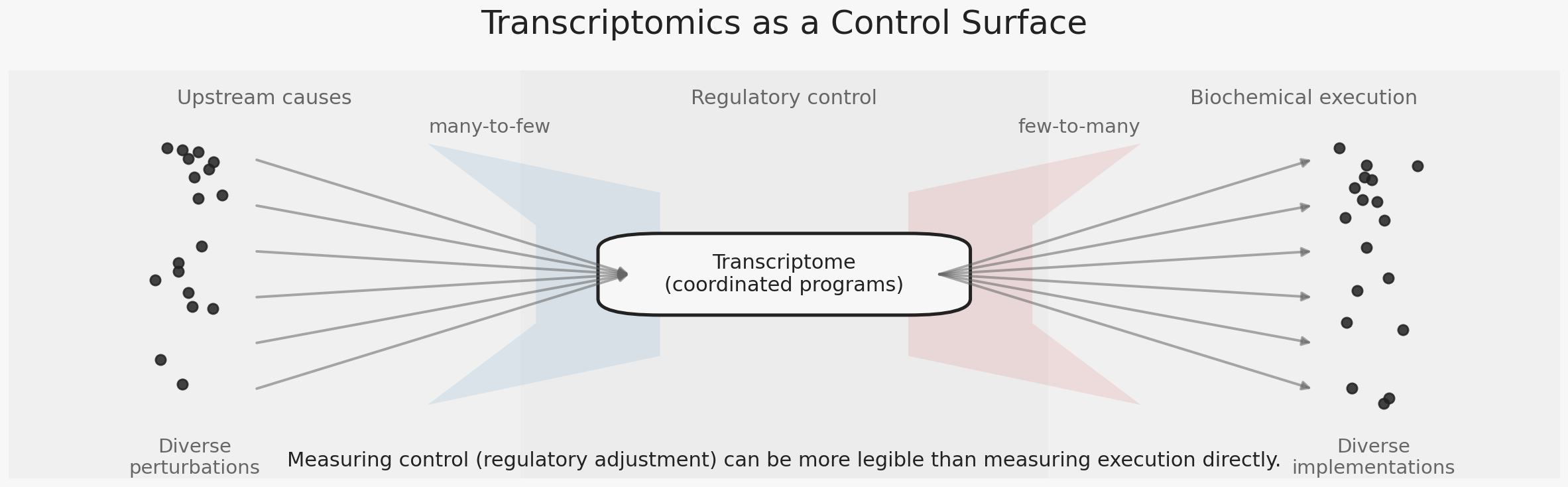
Transcription reflects how the cell reallocates resources and stabilizes itself under change. Many biochemical realities collapse onto a smaller set of regulatory adjustments — growth, stress adaptation, metabolic rewiring, inflammatory activation — producing patterns that remain statistically recoverable even in noisy data.
Different molecular stresses often converge on similar transcriptional responses. Disturbances in protein folding, metabolism, signaling, or structural integrity can all activate overlapping regulatory programs such as stress adaptation, inflammatory signaling, or growth arrest. This convergence is not accidental. Transcription functions as a control surface through which cells stabilize behavior despite diverse perturbations.
Why large datasets became interpretable
As sequencing scaled, the primary shift was statistical. Larger datasets increased the signal-to-noise ratio, making coordinated transcriptional programs easier to distinguish from technical variation. With enough observations, recurring patterns become separable from measurement noise, reinforcing the stability of regulatory structure across contexts.
Single-nucleus workflows extend this by trading molecular depth for robustness. They enable broader tissue access, frozen samples, and larger cohorts. The measurements are sparser per cell but more comparable across studies, strengthening the same regulatory structure that enables integration.
Perturbation within a constrained system
Pooled perturbation approaches coupled to transcriptomic readouts build directly on this property. Each cell carries a single indexed genetic perturbation, allowing many interventions to be evaluated simultaneously while preserving attribution. Responses organize along recognizable biological programs because perturbations act within an already regulated network.
This makes transcriptomics particularly well suited to settings where defined targets or pathways are being tested. Rather than searching for entirely new phenotypes, the assay measures how interventions redistribute coordinated programs.
What transcriptomics answers — and what it does not
RNA sequencing is a readout of regulation: what the system is attempting to do and which programs it engages. That is not the same as the realized biochemical condition of the cell, where protein activity, localization, complex formation, and post-translational control dominate.
Transcriptomics reports control, not execution. It captures how the system organizes its response, while many decisive differences are implemented downstream. The constraint is not on the biology itself, but on what can be inferred from regulation alone. Questions framed around coordinated response or pathway engagement map naturally to this layer.
This is one reason transcriptomics often performs well in target-driven and mechanistic settings. When the biological starting point is known, regulatory adjustment can be highly predictive of downstream behavior. In more open-ended contexts, functional differences mediated by protein activity, metabolism, or non-coding regulation may be only partially visible at the transcriptional level.
The point is not that one layer is superior to another. Each measurement system emphasizes different aspects of cellular organization. Transcriptomics is distinctive because it captures coordinated control, and therefore defines a particular kind of question that can be asked cleanly at scale.
Conclusion
Transcriptomics did not transform research simply by increasing dimensionality. It succeeded because it measures a layer of biology where regulation has already imposed order. Cells continually reshape transcription through signaling, metabolic feedback, and stress responses, producing coordinated programs that can be compared and modeled.
Its influence comes from this alignment between measurement and control. Transcriptomics reveals how biology attempts to stabilize itself — a perspective especially valuable when interventions are defined and the goal is to understand how regulated systems adjust, even when the biochemical consequences are more complex than transcription alone can reveal.