Biounfold #21 — Dynamic Experiment Design
Moving Beyond Static Assays
Most biological experiments are built around destructive measurements. We perturb cells, measure them once, and discard them. This approach has been powerful, but it limits our ability to study transitions and causal dynamics. Many cellular processes unfold over time, and single end-point measurements often mix primary and secondary effects. As a result, understanding how and when to observe cells is becoming as important as deciding what to measure.
Live-cell imaging changes this assumption. Because the readout is non-destructive, cells can be observed continuously. Instead of producing snapshots, experiments generate trajectories. This allows researchers to capture transitions, variability, and temporal patterns that are difficult to infer from static measurements. In practice, this transforms the structure of experiments. Cells can be monitored over time, and downstream assays can be triggered only when specific behaviors emerge. For example, one could follow morphological or signaling changes in real time and then apply sequencing or proteomics selectively to cells that exhibit a phenotype of interest. This increases signal, reduces cost, and focuses measurement on biologically relevant states.
Timing as a Design Variable
Timing becomes a central parameter in this framework. Early signals are often closer to causal mechanisms, but they are subtle and harder to detect. Later signals are easier to measure but increasingly confounded by adaptation, secondary effects, and stochastic variability. The longer an experiment runs, the more biological heterogeneity accumulates. As a result, experimental design involves balancing sensitivity and interpretability in time. Choosing when to observe cells can determine whether a signal is meaningful or obscured.
This is not only a biological problem but also a statistical one. Short observation windows reduce confounding but require stronger detection methods. Longer windows increase statistical power but reduce causal clarity. Advances in machine learning and computer vision are beginning to make it possible to quantify dynamic phenotypes at scale, which makes these timing trade-offs more explicit and measurable.
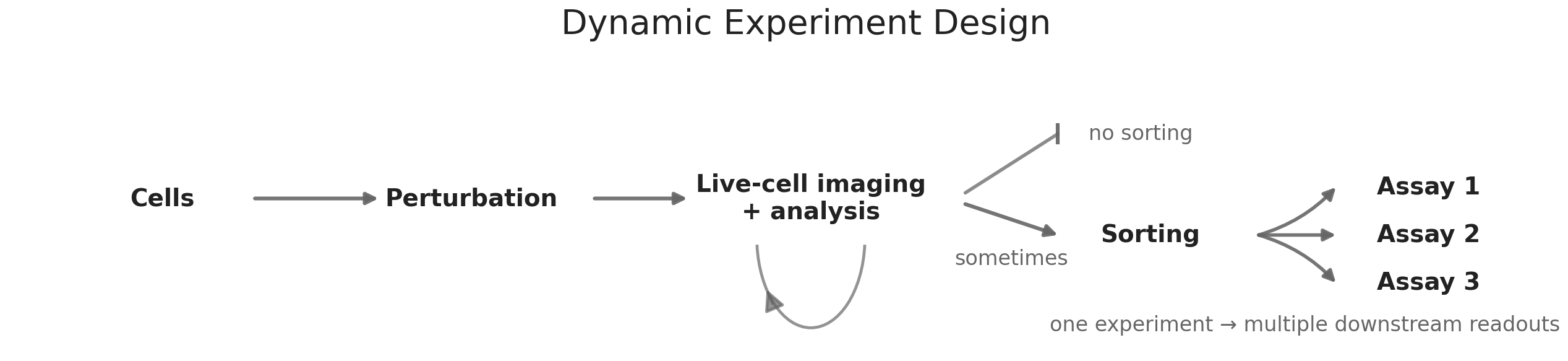
Cell sorting extends this logic by enabling active selection of cellular subpopulations based on observed signals. Techniques such as fluorescence-activated cell sorting allow populations to be divided either deterministically or probabilistically. When combined with live-cell imaging, sorting makes it possible to enrich for cells based on dynamic behavior rather than static markers. Subsets of cells can be directed toward different perturbations, environments, or downstream assays depending on their trajectory. In practice, enrichment often involves physically displacing cells that match specific temporal or phenotypic criteria, creating a bridge between observation and intervention.
Selective Measurement and Resource Allocation
A practical consequence of these developments is a shift in how experimental resources are allocated. Broad phenotypic readouts can be used to narrow the search space, allowing more expensive and mechanistically rich assays to be applied selectively. Some discovery platforms have already started to use imaging as an early prioritization layer. At Recursion, large-scale phenotypic imaging is used to identify perturbations and conditions that produce meaningful cellular responses before deeper molecular characterization. This tiered approach helps focus downstream assays on the most informative biological states rather than sampling broadly and uniformly.
A concrete example can be seen in immune cell activation. Early morphological and signaling changes during T cell engagement can be detected through live imaging, even when transcriptional changes remain subtle. By monitoring these early dynamics, it becomes possible to enrich for cells that show productive activation and then isolate these populations through sorting before applying molecular assays to characterize the underlying pathways. This reduces noise, improves statistical power, and concentrates effort on the most relevant states. This makes it possible to run faster and more focused cycles of experimentation.
For many years, the industry focused on standardizing around single assay types, especially high-throughput sequencing. This phase enabled large datasets and reproducible measurements, but it also highlighted the limitations of relying on one modality. As tools mature, integrating live-cell imaging, sorting, and molecular profiling is becoming increasingly practical. These approaches complement each other by capturing different aspects of cellular behavior and by enabling more flexible experimental strategies.
Together, live-cell imaging and sorting expand the range of what can be observed without destroying the system. They support experiments that emphasize dynamics, variability, and selective measurement. This shift is already influencing how discovery teams design experiments and where they invest their resources.