Biounfold #22 — Binding Affinity and the Geometry of Belief
Binding affinity is often framed as a technical objective. How tightly does a molecule bind? How much potency do we need? How low can the dose be? This framing is correct, but incomplete. Affinity does something deeper: it turns mechanistic beliefs into testable constraints.
The common story
In most drug discovery programs, affinity is treated as a lever of efficiency. Stronger binding enables lower doses. Lower exposure reduces off-target interactions and improves safety margins. This logic is sound and supported by decades of medicinal chemistry.
From this perspective, affinity is a parameter to optimize alongside solubility, permeability, clearance, and selectivity. Once nanomolar potency is reached, the program can move forward. Further gains may help, but diminishing returns are expected.
This view explains the appeal of structure prediction, docking, and generative chemistry. They promise faster access to molecules that bind. But this is only part of the story.
The deeper role of affinity
In practice, affinity does not only improve potency. It stabilizes hypotheses.
Weak and noisy signals keep multiple explanations alive. Targets remain ambiguous. Alternative mechanisms cannot be ruled out. Unexpected phenotypes remain difficult to interpret.
Strong and reproducible engagement changes this. It anchors the system. A mechanistic interpretation becomes dominant. Structure–activity relationships become interpretable. Confidence increases.
In this sense, affinity converts ambiguity into optimization. Discovery shifts from exploration to convergence. The landscape appears smoother. Progress becomes measurable. Decisions accelerate. This stabilizing effect is one of the main reasons affinity is so powerful. But it also introduces a structural risk.
The hidden fork
At some point, most programs encounter inconsistencies.
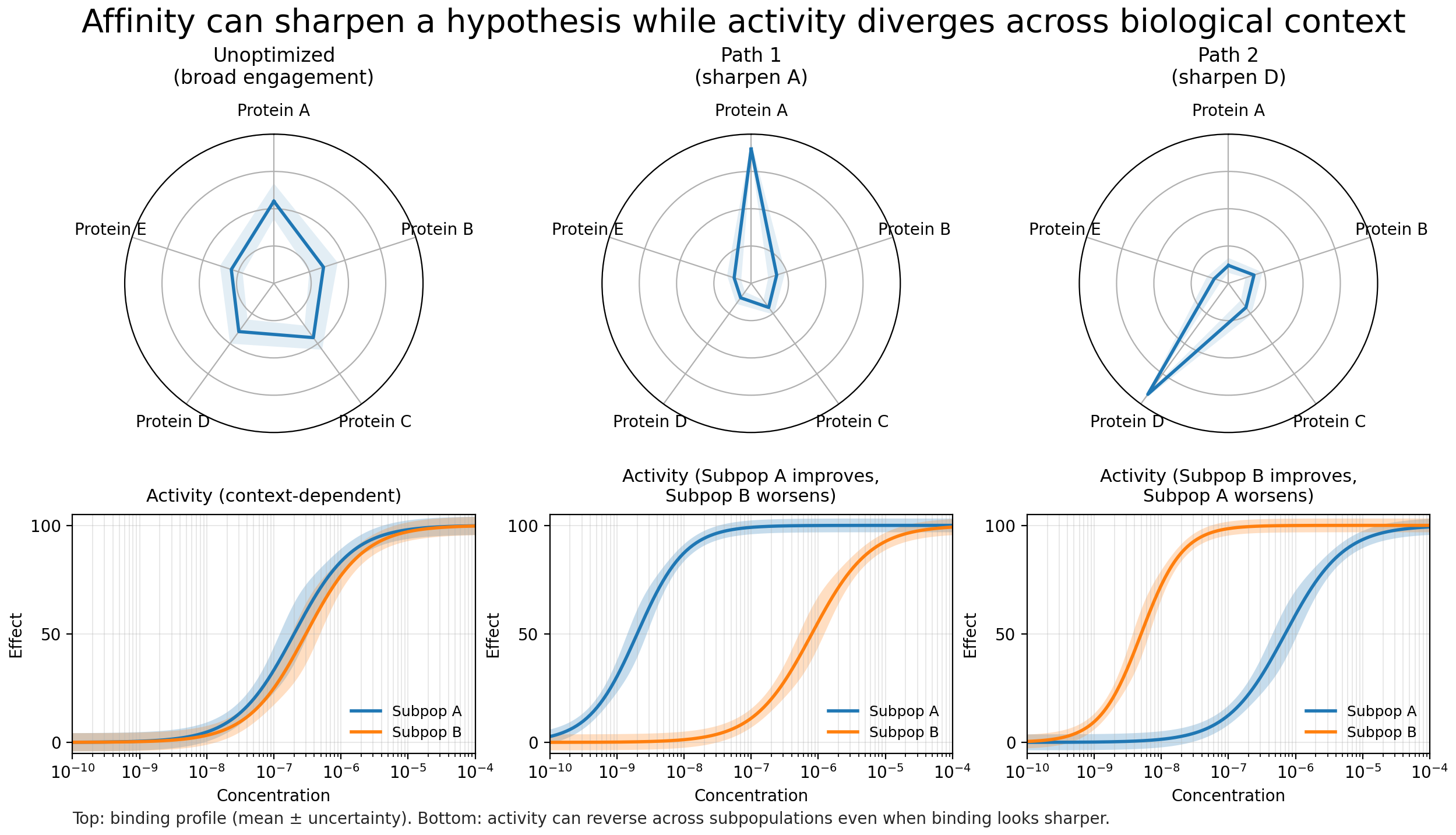
Two systems that should behave similarly do not. The same molecule produces different outcomes in related models. Unexpected phenotypes emerge. In heterogeneous diseases such as cancer, variability is expected. But persistent disagreement between models is a different signal. It suggests that the mechanistic framing is incomplete.
This is the moment when discovery reaches a fork. One path treats inconsistency as noise to manage. The other treats it as evidence that the hypothesis space remains wide. This decision is rarely explicit. It is embedded in timelines, resource allocation, and technical discussions. Yet it shapes the trajectory of the program.
The usual path: narrowing
The most common response is to narrow.
The patient population is redefined. Models that agree are prioritized. Biomarkers are refined. Heterogeneity becomes precision. This approach is rational and has produced real advances.
Strong affinity and clear engagement reinforce this direction. Signal becomes cleaner. Variability appears reduced. The mechanistic story stabilizes. The program gains speed and focus. But the hypothesis space also contracts. Alternative explanations receive less attention. Disagreeing models become secondary. Weak signals fade.
This trade-off is rarely discussed, but it is fundamental.
The alternative: treating inconsistency as discovery
A different path is possible.
Instead of resolving disagreement through narrowing, the inconsistency itself becomes the object of study. Why do these systems diverge? What biology is missing? Which assumptions are failing? This path is slower and more uncertain. It requires orthogonal evidence, multiple modalities, and tolerance for ambiguity. It often conflicts with time pressure and investment dynamics. Yet it can reveal hidden structure: new mechanisms, cryptic binding modes, resistance pathways, or previously unrecognized subpopulations.
Maintaining multiple chemical series is already a form of structured exploration. Diverse chemotypes probe the system through different interaction patterns and help expose hidden assumptions. But when biological consequences can be measured directly, they provide a stronger constraint on which hypotheses about mechanism and disease are actually valid.
Exploring biology makes these assumptions more explicit and measurable. Orthogonal systems, distinct cellular contexts, and alternative readouts do not replace chemical diversity. They extend it. In this sense, chemical and biological diversity are complementary ways of stress-testing the same causal model.
Platform implications
Affinity is not only a pharmacological property. It is a force that shapes the geometry of belief. It reduces uncertainty, but also redistributes it. It accelerates convergence, but can accelerate premature closure. It improves efficiency only when the abstraction is aligned with reality. Modern discovery platforms must therefore balance anchoring and breadth.
This requires:
- Designing loops where strong molecular hypotheses do not suppress orthogonal evidence.
- Maintaining diversity in both chemistry and biology.
- Treating disagreement between models as information rather than risk.
- Using affinity to stabilize learning without collapsing uncertainty too early.
Artificial intelligence is already shifting this balance. Most systems improve local optimization: predicting binding, proposing analogs, and expanding chemical space around existing hypotheses. This has dramatically increased the speed of convergence.
But faster convergence amplifies structural risk. If the dominant hypothesis is incomplete, optimization deepens commitment. The next generation of AI will likely broaden rather than only strengthen belief. It will integrate signals across modalities, highlight inconsistencies, propose alternative mechanisms, and suggest new experimental directions. AI will not only generate molecules. It will reshape the hypothesis landscape.
Binding affinity will remain central. But its deepest value is not that it makes drugs stronger. It is that it makes hypotheses testable. The question is not only how tightly our molecules bind. It is how tightly our assumptions are constrained.