BioUnfold #23 — ADME and the Biological Time Dimension
Drug discovery often begins with a clean picture. A molecule binds a target, the mechanism appears clear, cells respond as expected, and structure–activity relationships become interpretable. At this stage, molecules still behave like objects. They have properties such as potency, selectivity, or solubility. They can be compared, ranked, and optimized in relatively controlled systems.
ADME changes this perspective. Absorption, distribution, metabolism, and excretion are not only technical hurdles. They are the point where molecules stop being static objects and start becoming dynamic interventions unfolding in time and space.
From properties to trajectories
Binding and most cellular assays are conducted in environments where key variables are controlled. The molecule is present where we want it, at the concentration we decide, for the duration we choose. This isolation is what makes optimization possible.
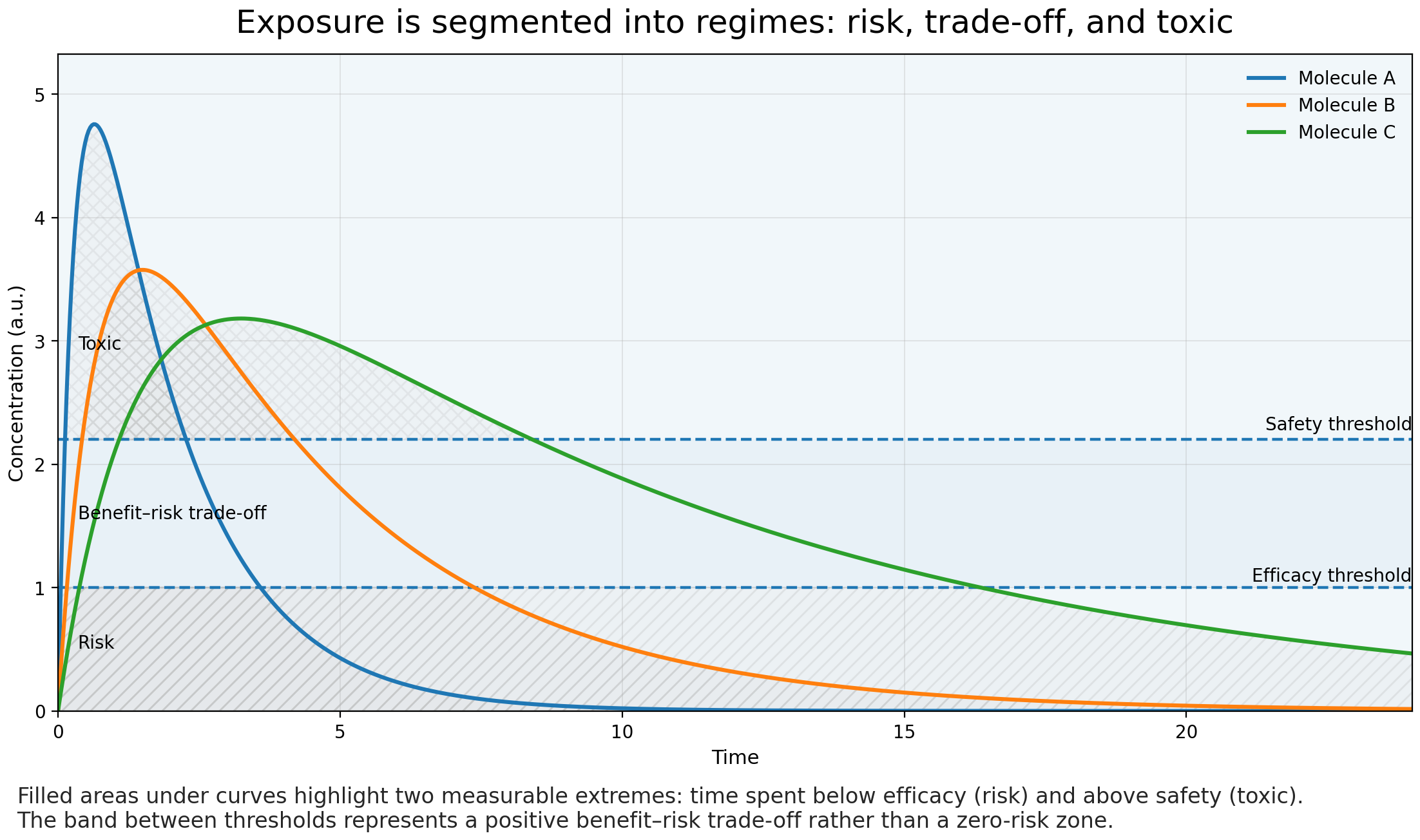
After administration, this control disappears. Molecules dissolve, cross barriers, accumulate in tissues, are transformed by enzymes, and eventually cleared. Exposure becomes dynamic. Two compounds with identical potency can produce very different biological outcomes simply because they reach the target differently over time.
Pharmacokinetics therefore acts as a translation layer between chemistry and biology. The central question shifts from what a molecule does to how its effects unfold over time in a living system.
ADME depends on mechanism
Because of its central role, ADME is often pushed earlier in programs. This reflects the real observation that many failures are driven by exposure rather than binding. However, without a mechanistic hypothesis, ADME data is difficult to interpret.
The optimal half-life, tissue distribution, or exposure level depends on how the biological process must be modulated. Some mechanisms require sustained engagement, while others benefit from transient or localized activity. Systemic exposure may be necessary in certain diseases and unnecessary or dangerous in others.
In this context, running ADME too early can generate large amounts of data that remain ambiguous. The same pharmacokinetic profile may be seen as favorable or problematic depending on the intended biological effect.
A progression in understanding
Most effective discovery programs move through a progression, even if it is not always explicit. They begin with an understanding of disease biology and build evidence that modulation of a specific process produces a meaningful phenotype. This may occur before or after the identification of a precise target, but both phenotype and mechanism must eventually be anchored.
Only then does ADME become strongly informative. Once the desired biological effect is defined, exposure can be evaluated in relation to that goal. The required duration, depth of engagement, and variability become concrete design objectives rather than abstract thresholds.
Why phenotype matters before ADME
Phenotypic validation plays a central role in this transition. If a molecule does not produce the expected biological effect in controlled systems, optimizing exposure rarely changes the outcome. Simplified assays do not capture the full complexity of organisms, but they remain powerful filters.
Orthogonal readouts strengthen confidence that the mechanism is correct and that the biological change is robust. They also provide early insight into how much modulation is required, how quickly effects emerge, and whether compensatory mechanisms appear. These observations directly shape ADME strategy.
This stage also influences decisions beyond the laboratory. Estimates of clinical benefit, acceptable risk, and potential patient populations depend on how convincing the early biological signal appears. Programs built on fragile or narrow readouts often face greater uncertainty later.
ADME as focused optimization
At this stage, ADME remains an optimization phase. Assays isolate individual dimensions of molecular behavior such as solubility, permeability, metabolic stability, or protein binding. The work is still iterative and controlled. The goal is to design molecules whose exposure patterns align with the intended biological mechanism.
What changes is the direction of optimization. Instead of treating ADME as a broad screen for generic liabilities, it becomes a structured effort to enable a specific therapeutic strategy. Trade-offs become explicit. Chemistry decisions become more intentional. Exposure is no longer abstract but directly connected to biological intent.
This makes ADME the last phase in which discovery retains strong experimental control before entering systems where effects become entangled and less separable.
Preparing for complex biology
By the time programs move into in vivo efficacy models, pharmacology, variability, and adaptation become tightly coupled. ADME helps reduce uncertainty before this transition by testing whether the desired biological modulation is physically achievable. It allows teams to reshape chemistry, modality, or dosing logic while iteration remains relatively fast.
This preparation is particularly important because many programs do not fail due to incorrect biology but because the required exposure profile is incompatible with safety or feasibility. These constraints are often visible earlier if exposure is evaluated in relation to mechanism.
Implications for AI and discovery platforms
Many of these ideas are already influencing modern discovery platforms. Integration of pharmacokinetics with mechanism, phenotypic readouts, and design is no longer theoretical. Multimodal datasets and exposure–response modeling are increasingly used to guide programs.
What remains challenging is not the existence of these components but their coordination. Mechanistic insight, phenotypic data, and pharmacokinetics are often generated in parallel but interpreted separately. The opportunity is to treat them as parts of a single learning system that evolves as biological understanding deepens.
In practice, this means connecting experimental loops rather than multiplying them. It also means designing data generation around decision points rather than around assay availability.
Conclusion
ADME is often described as a technical domain within drug discovery. In practice, it is a phase where optimization becomes grounded in biological intent. Once disease biology, phenotype, and mechanism are sufficiently anchored, exposure becomes a design variable rather than a generic constraint.
Binding stabilizes the hypothesis. Phenotype confirms that the biology matters. Mechanism clarifies how modulation produces benefit. ADME aligns molecular behavior with that objective.
This alignment allows discovery programs to enter more complex biological systems with greater clarity about what success should look like and why.