BioUnfold #24 — Toxicity: When Discovery Meets Consequence
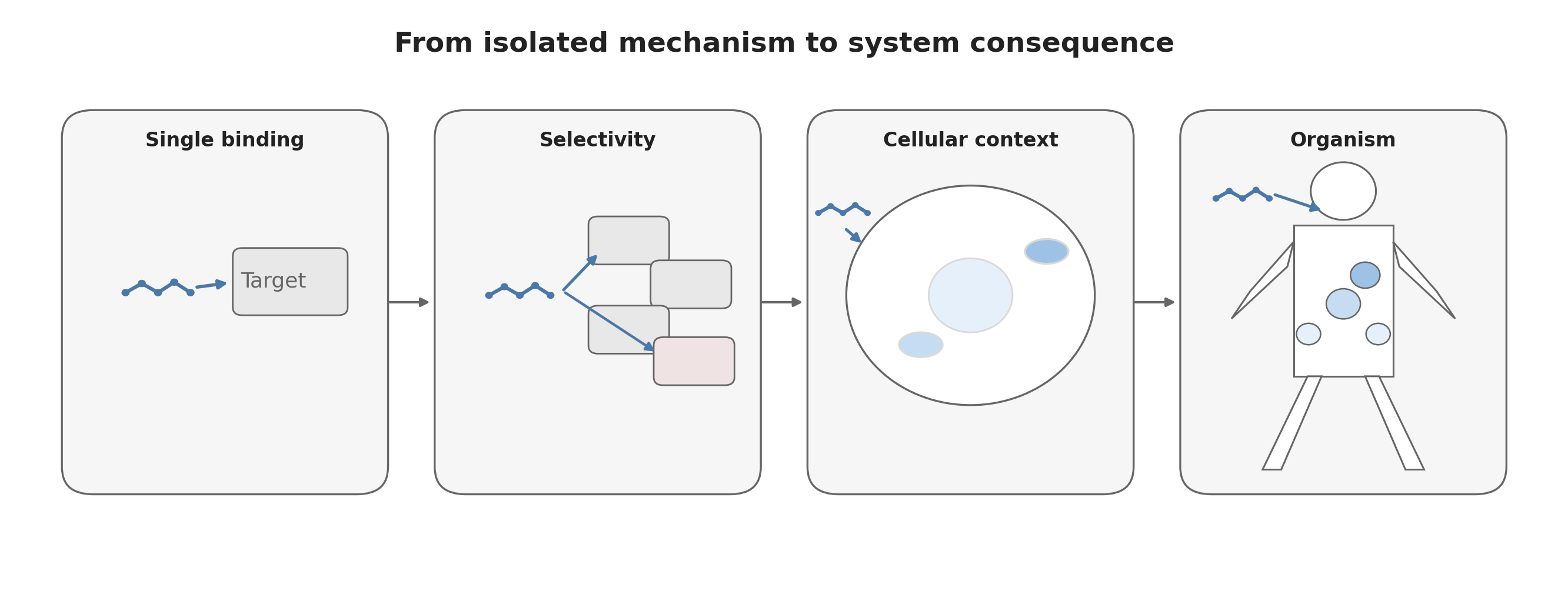
Drug discovery is built on reductionism. Proteins are isolated, pathways are studied individually, and molecules are optimized against specific targets. This approach works remarkably well. It allows scientists to understand biological mechanisms, design selective molecules, and progressively refine chemical matter.
Toxicity appears when the molecule leaves these controlled systems and enters the full organism.
Unlike potency or exposure, toxicity is not a single property. It is the manifestation of unintended biological consequences unfolding across time, tissues, and physiological contexts.
Many forms of toxicity
Toxic effects rarely follow a single pattern. They can emerge along several dimensions and timescales.
Some appear rapidly after exposure, reflecting direct disruption of essential cellular processes. Others develop slowly as subtle damage accumulates over weeks or months. Certain toxicities are tissue-specific. The liver, kidney, and heart are frequent sites because they concentrate drugs, metabolize them, or rely on tightly regulated physiological processes.
In other cases, toxicity depends strongly on context. Metabolic state, immune activation, or co-medications can all change how a molecule interacts with biology. Even within the same organism, different cell types may respond differently to the same compound. A drug that produces the desired modulation in one population may disrupt critical functions in another.
This diversity is what makes toxicity difficult to anticipate. It emerges from the interaction between molecular binding and the broader biological system.
Binding remains the clearest explanation
The most reliable way to understand toxicity is still to ask a simple question: where else does the drug bind?
Off-target binding explains many adverse effects. Ion channels, transporters, and enzymes involved in fundamental cellular processes are common sources of unexpected liabilities. Even relatively weak interactions can become relevant when exposure is high or sustained.
Understanding these interactions has improved significantly over the past decades. Advances in structural biology, target characterization, and chemical selectivity profiling now allow teams to identify many risks earlier in discovery. Programs today often begin with deeper knowledge of the intended target and its physiological context, which reduces the likelihood of unexpected toxicity emerging later.
However, binding alone is not always sufficient to explain toxic effects.
Biology amplifies small perturbations
Biological systems are highly interconnected. Modulating one protein can propagate through signaling pathways, metabolic networks, and tissue-level physiology.
A drug may bind its intended target with high specificity and still produce unintended consequences because the pathway it perturbs is linked to other essential processes. In these situations toxicity does not arise from off-target binding, but from on-target effects occurring in the wrong biological context.
This distinction is important because the mitigation strategies differ. Improving selectivity may not resolve a toxicity driven by the intended mechanism itself. Understanding the broader biological network therefore becomes as important as measuring molecular binding.
Expanding the search for hidden interactions
Because unexpected liabilities often originate from unknown interactions, discovery teams increasingly explore broader profiling strategies.
Large panels of biochemical assays, cellular systems, or tissue lysates can reveal previously unrecognized binding partners. Proteomics-based approaches, chemoproteomics, and phenotypic profiling can also expose hidden interactions between molecules and biological systems.
These strategies rarely eliminate toxicity altogether, but they help uncover potential mechanisms earlier in discovery. When unexpected biology appears, teams can decide whether to redesign the chemistry, adjust exposure, or reconsider the therapeutic hypothesis. In practice, these methods function less as simple filters and more as sources of mechanistic clues.
The limits of animal models
Animal studies remain a central step in evaluating toxicity, yet their predictive power is often misunderstood.
Many toxic effects observed in humans cannot be reproduced in mice or other preclinical species because the underlying biology differs. Enzyme expression, immune responses, and metabolic pathways can vary substantially across species.
For this reason, improvements in toxicity prediction over the past decades are not primarily the result of better animal models. They stem more from improved target understanding, molecular design, and earlier profiling strategies. Better biology upstream often prevents problematic molecules from progressing far into development.
Toxicity as a design constraint
In practice, toxicity becomes a constraint that shapes discovery decisions rather than a separate stage that follows them.
Chemistry teams adjust molecular structures to reduce off-target interactions. Exposure profiles are tuned through ADME optimization, and dosing strategies are adapted to avoid sustained perturbation of sensitive pathways.
Sometimes the conclusion is that the target itself is not therapeutically viable. In other cases the mechanism remains sound but requires a different modality or delivery strategy. Toxicity therefore acts as a feedback signal that forces the program to reconcile molecular design with physiological reality.
Implications for discovery platforms
Modern discovery platforms increasingly integrate toxicity signals earlier in the learning cycle.
Large selectivity panels, cellular profiling, and multimodal datasets provide richer descriptions of how molecules interact with biological systems. Machine learning models can help detect patterns across these datasets and highlight potential liabilities before they appear in vivo.
Yet the challenge is rarely the absence of data. It is the interpretation of complex biological signals that do not have simple explanations. Toxicity sits at the intersection of chemistry, biology, and physiology, and understanding it requires connecting these domains rather than analyzing each independently.
Conclusion
Toxicity is sometimes framed as the enemy of drug discovery. In reality it is the biological system revealing the full consequences of molecular intervention.
Binding explains intention. ADME determines exposure. Toxicity reveals consequence.
Programs that understand this progression can treat toxicity not only as a risk to avoid but as information about how biology truly responds to perturbation. Toxicity is therefore not only a safety problem. It is the moment when a carefully engineered molecular intervention encounters the full complexity of the organism.