BioUnfold #28 — Plate-wide Chemistry: From Hits to Evidence
Testing whether a signal survives expansion
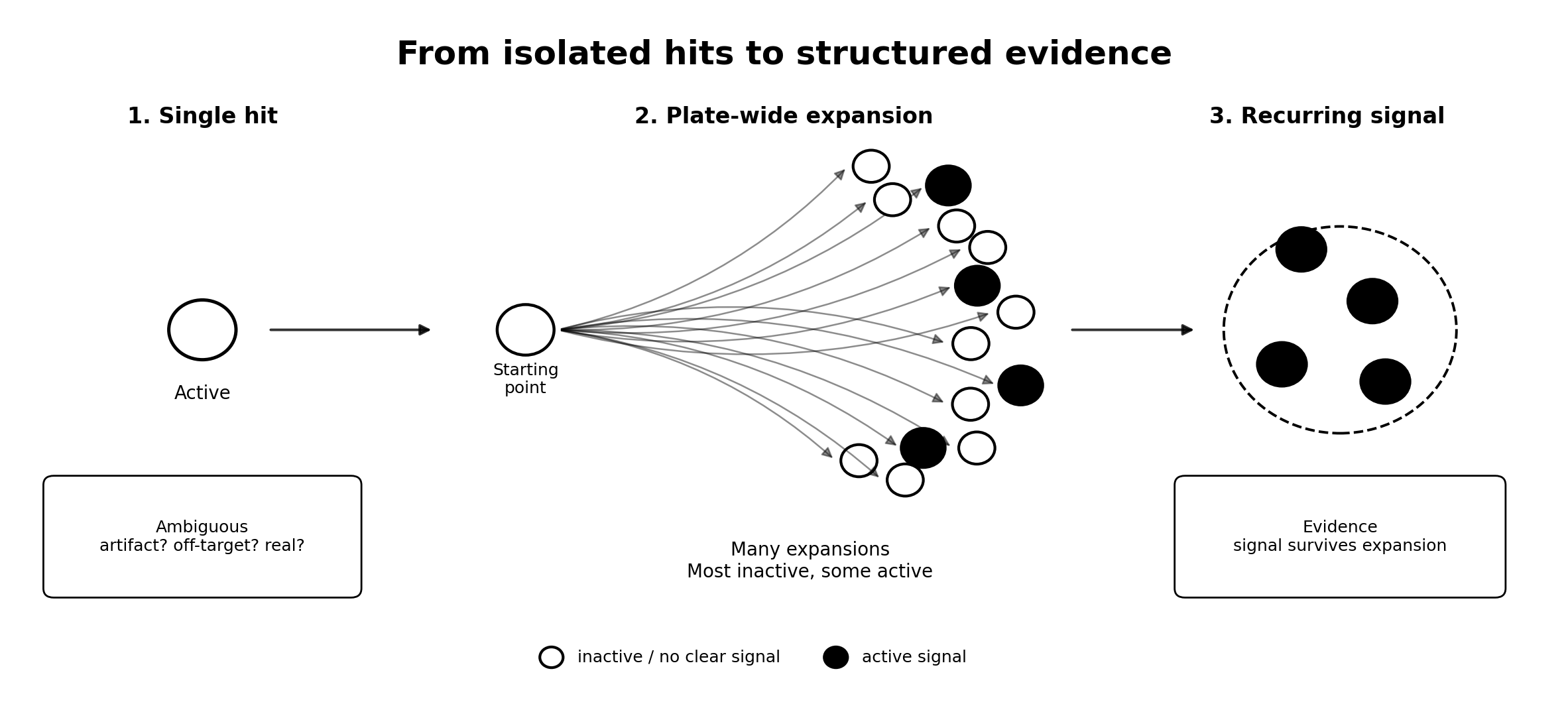
Most discovery workflows treat molecules as isolated decisions.
A compound is designed, synthesized, tested, and evaluated on its own. If it shows activity, it becomes the starting point for the next round. This approach is precise and has defined medicinal chemistry for decades. It works particularly well when structure–activity relationships are already understood and optimization is the goal.
But early in discovery, signals are fragile. A single hit is difficult to interpret. Activity may come from the intended biology, from off-target effects, from reactivity, or from assay artifacts. At this stage, the question is not yet how to optimize a molecule, but whether the signal itself is real enough to pursue.
Plate-wide chemistry changes how this question is approached.
Expanding instead of optimizing
Instead of designing and synthesizing one compound at a time, plate-wide chemistry generates many compounds in parallel. Starting from a weak hit or fragment, a set of transformations is applied across a plate, producing dozens to hundreds of molecules in a single round.
Rather than committing to a specific molecule, the experiment explores multiple directions in chemical space at once. Each compound represents one possible expansion of the starting point. Together, they test whether the initial signal can be reproduced under variation.
In practice, this comes with constraints. Reactions must be compatible across the plate, conditions must remain stable, and not all wells will produce usable compounds. Plate-wide chemistry trades flexibility for parallelism. It does not cover all of chemical space. It samples what is accessible under a given transformation.
Not optimization, but expansion
The molecules produced in a plate are often substantially different from the starting fragment. They are larger, explore different interactions, and do not form a smooth chemical neighborhood. As a result, classical structure–activity relationships rarely emerge at this stage.
Activity does not vary continuously. Many compounds will be inactive. Some will show signal. The goal is not to observe gradients, but to observe recurrence.
What matters is whether activity appears repeatedly across independent compounds derived from the same starting point. A single active molecule remains ambiguous. Multiple active molecules suggest that the signal is not accidental.
Plate-wide chemistry therefore does not validate individual compounds. It tests whether a signal survives expansion.
From hits to evidence
This shift is subtle but important.
In a sequential workflow, a hit is evaluated on its own. Confidence builds slowly through iterative design. In a plate-based workflow, confidence emerges from patterns across compounds.
A fragment that produces one active molecule may not be meaningful. A fragment that produces several active molecules, even if structurally diverse, carries more weight. The signal has survived variation.
The unit of evidence is no longer a single compound. It is a set.
This is particularly useful in phenotypic discovery, where mechanism is unknown and signals are harder to interpret. Plate-wide chemistry does not resolve the underlying biology, but it helps determine whether there is enough consistency to justify deeper investigation.
Between exploration and optimization
Plate-wide chemistry occupies a specific place in the discovery process.
In many screening campaigns, libraries already contain clusters of related compounds, and early signals of consistency can emerge directly from primary data. Plate-wide chemistry becomes most useful when such structure is missing — when starting from fragments, novel chemotypes, or sparse hits where analogs are not readily available. In these cases, generating controlled variation around a starting point does not only support human interpretation. It also creates consistent datasets that can be used to guide subsequent design, whether by chemists or by learning-based systems.
Upstream approaches such as DNA-encoded libraries explore large regions of chemical space but provide limited resolution. Downstream medicinal chemistry focuses on precise optimization but requires a reliable starting point.
Plate-wide chemistry sits between these regimes. It takes uncertain starting points and tests whether they can generate consistent signal when expanded. It increases resolution without requiring full optimization. It is therefore most valuable early, when decisions are about direction rather than refinement.
Constraints and trade-offs
The advantages of plate-wide chemistry come with clear limitations.
Reactions must be robust and compatible across many compounds. Byproducts and incomplete reactions can complicate interpretation, requiring analytical validation. Not all chemical transformations are accessible in this format, and the resulting molecules may not cover all relevant design hypotheses.
Throughput does not scale indefinitely. Running multiple plates introduces logistical complexity, and the benefits depend on maintaining a high proportion of usable compounds.
Most importantly, plate-wide chemistry does not replace medicinal chemistry. It does not produce optimized molecules. It produces evidence that guides where optimization should begin.
Testing whether a signal is real
Early discovery is often framed as a search for active compounds. In practice, it is a search for signals that can be trusted.
Plate-wide chemistry reframes this search. It does not make individual results more certain. It evaluates whether they remain consistent under variation.
A single hit is fragile. A recurring signal is not.
Plate-wide chemistry is not about making many compounds.
It is about turning isolated hits into structured evidence.